Disclaimer: Think carefully before exposing your data to any third party — this is doubly important with health data. Everything here is for fun and in no way should be construed as medical advice.
Overview
A few years back I was diagnosed with type 2 diabetes — due to a series of lab errors. This BBC News article has more detail, but in short: my HbA1c was reported five points over its actual value, making me officially diabetic. It came as a shock, and brought with it a whole lot of worry.
Shortly after I got the news, I was invited to the local surgery to have a Libre 2 sensor explained and fitted. These are small discs that stick a needle into your arm and report your blood sugar to an app on your phone.
Abbott’s LibreView dashboard is fine for glancing at trends. But I wanted to interrogate my data: correlate glucose spikes with meals, compare patterns across weeks, ask questions I hadn’t thought of yet. Most of all, I wanted my data on my server.
This post covers step one: building an MCP server to make that data conversational.
What is MCP?
Model Context Protocol is Anthropic’s open standard for connecting AI assistants to external data sources. Instead of copying and pasting data into Claude, you build a small server that exposes tools Claude can call directly.
You define what data is accessible and how it authenticates. Claude does the rest.
For my glucose data, this means I can ask things like:
- “What was my average blood sugar last week?”
- “Show me a graph of my readings over the past 24 hours”
- “What time of day do I typically see spikes?”
And Claude will fetch live data, analyse it, and respond conversationally.
The Architecture
Before the code, a few design decisions worth explaining.
FastMCP over raw MCP: The mcp library has a low-level server API, but FastMCP gives you a decorator-based interface that’s much closer to how you’d write a Flask or FastAPI app. For a project this size, the abstractions are net positive — you get tool registration, schema generation, and transport handling for free.
stdio transport: MCP supports both stdio and HTTP transports. For a local desktop integration (Claude Desktop), stdio is simpler — no ports, no auth, no networking. The server process is spawned by the client and communicates over stdin/stdout. The trade-off is that it’s desktop-only; an HTTP transport would be needed for remote access.
uv for dependency management: Using uv keeps the project isolated and makes the Claude Desktop config clean. The entire run command is uv --directory /path/to/gcm-mcp run main.py — no virtualenv activation, no path juggling.
Connection pooling: One thing to be aware of with the pattern below is that you don’t want to re-authenticate on every tool call. In the real implementation, the PyLibreLinkUp client is initialised once at module load, not inside each function.
The Implementation
The server is around 100 lines of Python. The core structure:
from mcp.server.fastmcp import FastMCP
from pylibrelinkup import PyLibreLinkUp
import os
mcp = FastMCP("blood_sugar")
# Initialise once at module load — not per-call
client = PyLibreLinkUp(
username=os.getenv("API_USERNAME"),
password=os.getenv("API_PASSWORD")
)
@mcp.tool()
def get_blood_sugar() -> dict:
"""Fetch current blood sugar reading from Libre sensor."""
return client.get_current()
@mcp.tool()
def get_blood_sugar_history() -> dict:
"""Fetch blood sugar history from Libre sensor."""
return client.get_history()
if __name__ == "__main__":
mcp.run(transport="stdio")The @mcp.tool() decorator handles everything: registering the function, generating a JSON schema from the type hints and docstring, and exposing it to Claude. When you ask “what’s my current blood sugar?”, Claude matches the question to the right tool, calls it, receives the JSON, and formulates a human-readable reply.
Configuration lives in Claude’s desktop config file. usually ~/Library/Application\ Support/Claude/claude_desktop_config.json:
"mcpServers": {
"blood_sugar": {
"command": "uv",
"args": ["--directory", "/path/to/git-checkout/gcm-mcp", "run", "main.py"],
"env": {
"API_USERNAME": "your@email.com",
"API_PASSWORD": "your-password"
}
}
}What Can You Do With It?
Simple queries: “What’s my blood sugar right now?” returns the current reading with trend direction.
Analysis: “Draw a graph of my blood sugar over the past 24 hours” – Claude fetches the history, generates a visualisation, and displays it inline.
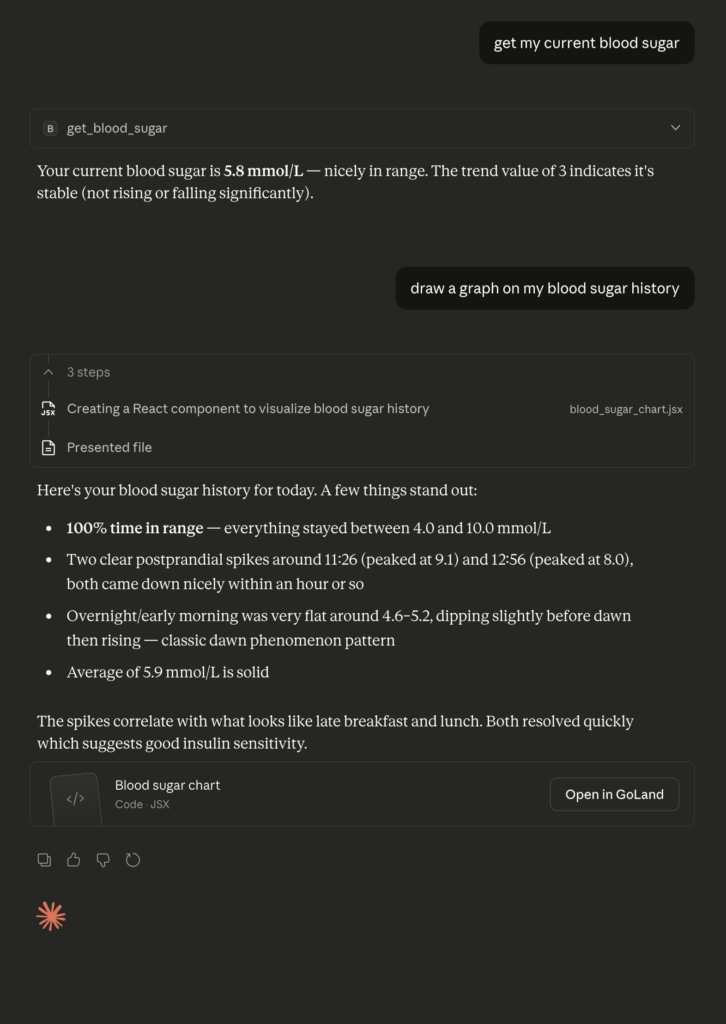
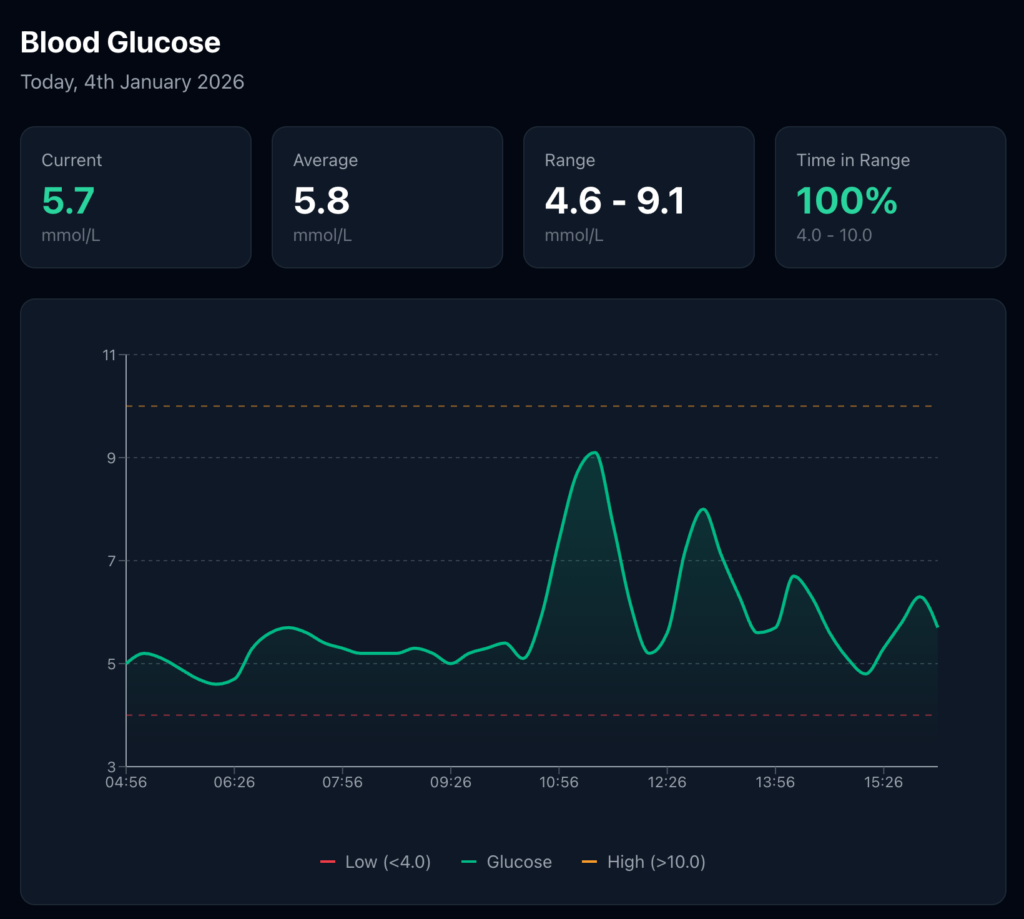
Pattern recognition: “When do I typically see my highest readings?” – Claude analyses the timestamps and surfaces patterns I might have missed in a static dashboard.
The key insight is that this isn’t just a dashboard replacement. It’s a conversational interface to my health data. I can ask follow-up questions, request different visualisations, and explore correlations – without switching contexts or learning a new tool.
Why Build This?
Partly curiosity. MCP was new and I wanted to understand how it worked from the inside.
Partly practical. I genuinely wanted better access to my glucose data, and the official tools weren’t designed for interrogation.
But mostly, I think something interesting is happening here. We’re moving from “apps that display your data” to “assistants that understand your data”. The interface is no longer a dashboard.
For health data, this matters. I don’t always know what questions to ask. A dashboard shows me what its designers anticipated. A conversational interface can explore the unexpected.
Try It Yourself
Try It Yourself
The code is on GitHub: github.com/chrishannam/gcm-mcp
You’ll need:
- A Freestyle Libre 2 Plus sensor (or compatible)
- A LibreLinkUp account connected to your sensor
- Claude Desktop with MCP support
- Python and
uvfor dependency management
Setup takes about ten minutes. The README walks through the configuration. Some Python knowledge is needed at the moment – there’s no GUI setup flow yet.
What I Built Next
After getting this working, I extended the same pattern to pull in other health data sources: blood pressure and body composition from my Withings devices, and activity data from Strava. I also added observability with Logfire so I could trace tool calls across all three MCP servers.
That architecture – multiple data sources, a unified conversational interface, and proper instrumentation – is covered in the next post: Building a Personal Health Data Platform with MCP, Withings, Strava, and Logfire (coming soon).
If you’re sitting on interesting personal data and want to make it conversational, MCP is worth exploring. Just take care, it’s your data, and no AI assistant is a substitute for a trained medical professional.
Chris Hannam is a software engineer with 20+ years of Python experience, currently based in Northern England. He’s interested in health tech, data engineering, and building tools that make complex information accessible.
Chris's Blog
My thoughts and projects.
Twenty Twenty-Five
Designed with WordPress